Steven Potiris has successfully completed his PhD thesis, entitled “Robust and Efficient Individual Plant Mapping of Vegetable Crops”. The work was supervised by Professor Salah Sukkarieh and Dr Asher Bender.
Congratulations Steven!
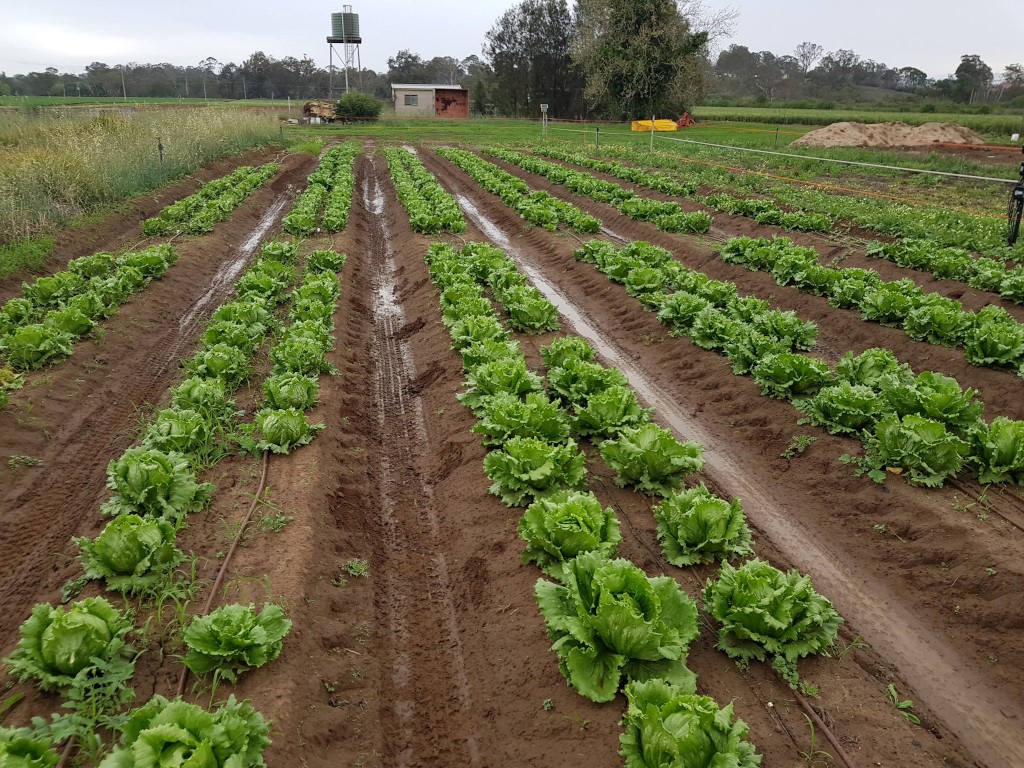
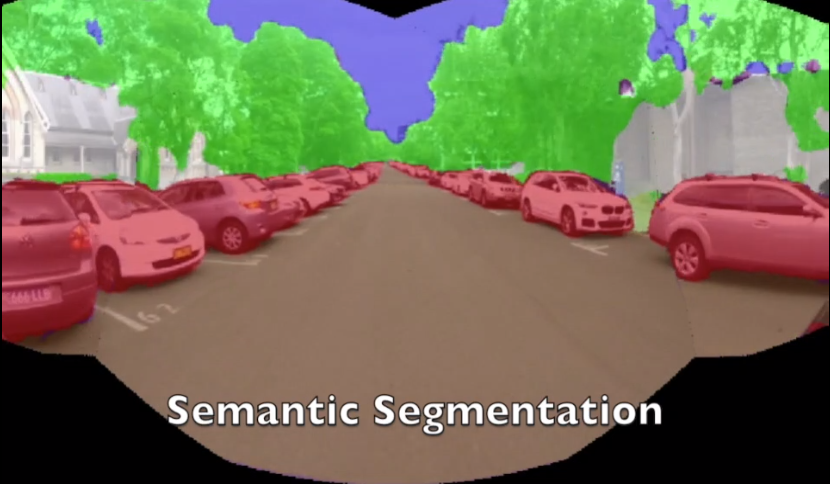
Crop mapping can provide phenotype data for breeding research. To provide phenotyping data for individual plants, a system must be able to detect and segment each plant from its background. Many existing datasets reflect non-commercial cases where the plants do not overlap with weeds or other plants. The relative ease of segmenting the crops from background in these environments warrants the use of implicit instance segmentation models, which fail in more complex environments.
This thesis investigates the use of explicit instance segmentation models to overcome the shortcomings of implicit models. The aim is to enable an unmanned ground vehicle to produce spatio-temporal crop maps which attribute phenotype measurements from an RGB camera to individual plants, while being robust to clutter and overlap from both neighbouring plants and weeds.
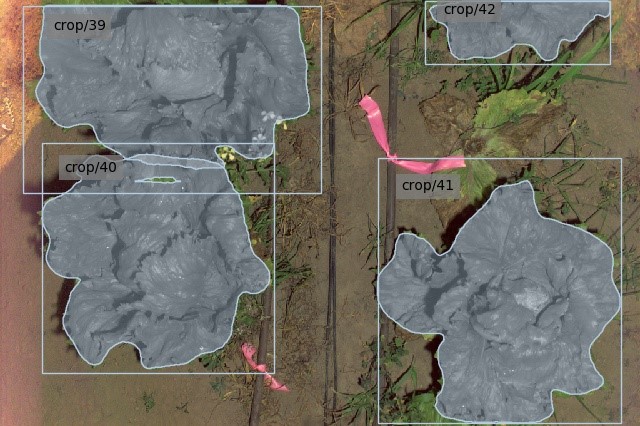
This thesis details the design and capture of a commercially focused dataset which allows overlap between adjacent crops and weeds. Various inputs (water, herbicides and nitrogen) have been applied in an experiment to create a diverse dataset among cos and iceberg lettuce, broccoli and cauliflower crops. The datasets are labelled and used to train both implicit and explicit instance segmentation models, and the results are compared against crop-crop and crop-weed overlap factors.
The best performing model is used in a crop mapping system where Ladybird, a high-throughput phenotyping platform, is used to calculate and spatio-temporally map phenotypes of individual plants in a field. The maps show a spatially accurate temporal representation of various phenotypes calculated from the RGB imagery obtained by Ladybird. The final individual crop maps are compared against accurate ground-truth maps, and the performance of the system as a whole is evaluated.